Deep Learning Model Boosts LC-MS Proteome Coverage with Alternative Techniques
Recent advancements in mass spectrometry have significantly enhanced our understanding of complex peptide mixtures. Utilizing the Omnitrap LC-MS platform, researchers developed innovative methods, including ultraviolet photodissociation (UVPD), electron transfer dissociation (ECD), and electron-induced dissociation (EID). These developments aimed to increase proteome coverage and improve peptide identification rates.
Enhancing Proteomics with Advanced LC-MS Techniques
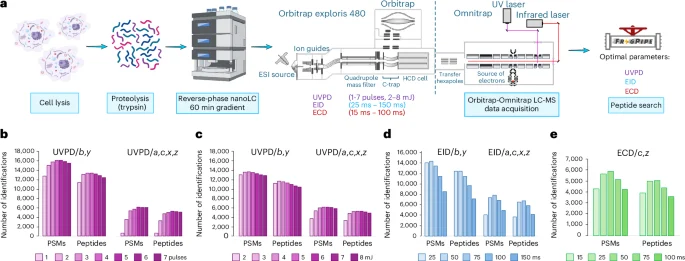
The Omnitrap system allows for the effective analysis of peptides through various dissociation techniques. Optimizing parameters such as laser pulses and energy levels is crucial for maximizing identification rates. The research demonstrated that four laser pulses at 6 mJ provided the best peptide spectrum matches (PSMs) for UVPD, while EID and ECD exhibited optimal identification rates at 50 ms of irradiation time.
Investigation of Fragmentation Techniques
- UVPD: Achieves maximum PSMs with four laser pulses at 6 mJ.
- EID: Optimal identification occurs at 50 ms irradiation.
- ECD: Also performs best with 50 ms using c and z fragments.
Further experiments diversified peptide sequences using multiple proteases, including trypsin and chymotrypsin, coupled with offline fractionation techniques. The combination of these methods significantly increased the depth of peptide analysis across various dissociation techniques.
Comparative Analysis of Fragmentation Techniques
In totals reported across five proteases, the results showed that:
- UVPD yielded approximately 1,141,000 PSMs.
- ECD followed closely with 1,070,000 PSMs.
- EID achieved around 900,000 PSMs.
- HCD, with a maximum efficiency, reported 1,160,000 PSMs.
Interestingly, while EID had the least identified PSMs, its acquisition speed of approximately 6.3 MS2 scans per second indicates its potential for efficient peptide sequencing.
Deep Learning Model for Fragment Intensity Prediction
A significant innovation introduced in this research is a deep learning model aimed at predicting fragment ion intensities for UVPD, EID, and ECD methods. This model was developed using comprehensive datasets generated during experimentation. By predicting fragment intensities, researchers can enhance the accuracy of peptide identifications, leading to higher success rates in LC-MS analyses.
Model Training and Validation
The deep learning model showed promising correlation metrics, with median Pearson correlations exhibiting a strong relationship between predicted and experimental values. This indicates high reliability in predicting ion intensities, which can greatly assist in improving database search results for peptide identifications from complex samples.
Impact on Database Search Efficiency
Utilizing predicted fragment intensities during database rescoring demonstrated remarkable improvements in peptide identification rates across multiple fragmentation techniques. By enhancing the identification ratio of observed ions to theoretically possible fragments, researchers achieved a clear separation between true-positive and decoy matches. This led to a significant increase in the recovery of estimated true positives at stringent FDR thresholds.
Key Findings from Rescoring Analysis
- The combination of intensity prediction with database searching enhanced identifications by 3% to 40.5% across different enzymes and techniques.
- Chymotrypsin and UVPD showed marked gains, while ECD maintained a stable identification rate due to its cleaner spectrum profile.
These advancements underline the transformative potential of deep learning in proteomics, paving the way for increased efficiency and accuracy in the analysis of complex biological samples. The ongoing integration of innovative methodologies stands to further enhance proteomic coverage and the identification of peptides, informing better biological insights.





